For the past few decades, the field of molecular evolution has largely focused on the evolution of individual gene families and overall genome structure. Analogous to the transition from reductionist to system-scale approaches in molecular biology, the logical next step in molecular evolution research is to study the evolution of genes in the context of the systems in which they function. One way to study how systems of interacting components evolve is to simulate the evolution of suitably abstracted system models in silico. Given recent developments in the field of high-performance computing, it is now possible to simulate the evolution of molecular systems at an unprecedented level of mechanistic detail. We use mathematical models to map the genotype of artificial transcriptional regulatory systems to their expression phenotype. We then use these models in population-scale evolutionary simulations to investigate how transcriptional systems evolve.
-
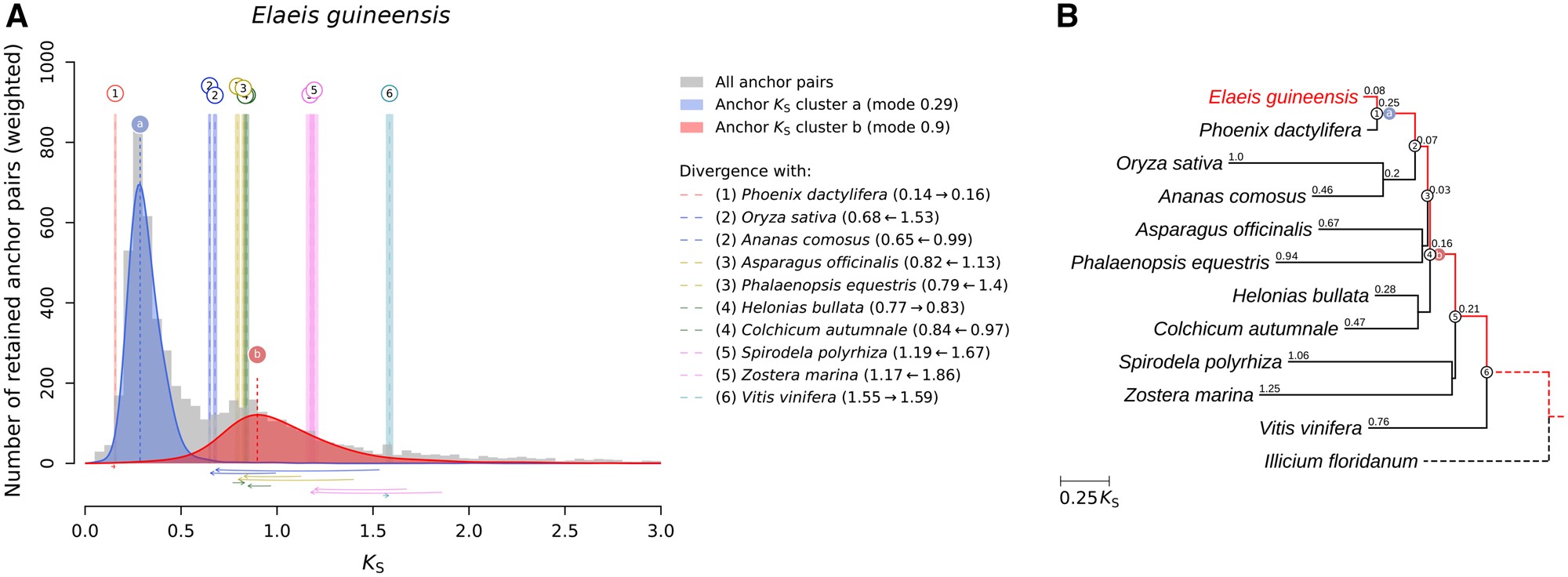 Modeling the evolution of transcriptional systems in silico
Modeling the evolution of transcriptional systems in silico